The naive Bayes classifier is a simple but effective classification algorithm which can be used for image segmentation/clustering. We have created a custom naive Bayes classifier and integrated it into the QCPA framework. This method is suitable only when you have prior knowledge of the maximum number of image segments expected, and their average colours, unlike the RNL Clustering technique. The means and standard deviations of each cluster across n-dimensions of channels in the image must be provided.
Video Guide
Input Image
This process requires a 32-bit image stack, such as a calibrated (linear & normalised) multispectral image stack, or a cone-catch image. It also requires a results table open which has the mean and standard deviations of each cluster for each image channel. The columns must be labelled with the channel name followed by “_mean” and “_SD” for means and standard deviations respectively. These results can easily be created by selecting one ROI as an example of each cluster in a multispectral image, and pressing “R” on the keyboard to provide means and standard deviations for each ROI. e.g. in the example below, a small section of “red” moth wing was selected as the first ROI, a small section of “black” for the black wing parts etc…
Running Naive Bayes Clustering
With the input image highlighted run:
plugins > micaToolbox > QCPA > Naive Bayes Quantisation
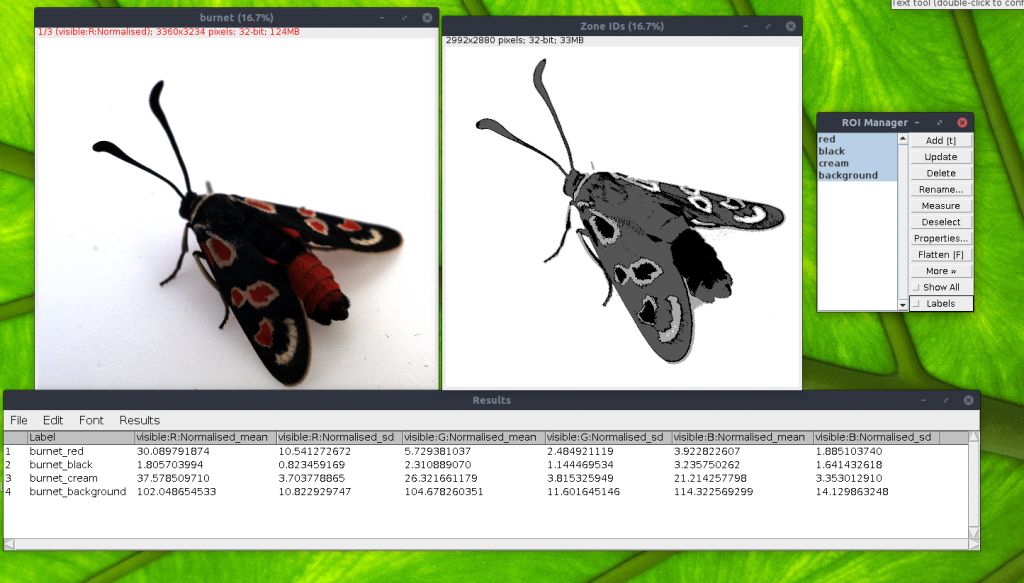
Output
The script outputs an image where each pixel value specifies a cluster ID. In the above example, the first measurement is “red” on the burnet moth wing, so parts of the image classified as “red” will have a pixel value of 1. “black” will be 2, etc… up to the full number of clusters (it can deal with up to 255 clusters, but if you have that many you should probably be using a different method!)
The output clusters can be converted into ROIs by using the “Cluster Particle Analysis” tool:
plugins > micaToolbox > QCPA > Cluster Particle Analysis
